GenElute™ 哺乳动物基因组 DNA 微量提取试剂盒实验方案 ;
产品描述
GenElute™哺乳动物基因组 DNA 微量提取试剂盒提供了一种从多种培养细胞、组织(包括啮齿动物尾巴)和新鲜全血或白细胞中分离纯基因组 DNA 的简单方便的方法。该试剂盒结合了二氧化硅和微型离心机的优势,无需使用昂贵的树脂、乙醇沉淀以及以及苯酚、氯仿等有害有机化合物。起始原料在含有离液盐的溶液中裂解,以确保大分子彻底变性。当裂解物在微量离心管中通过二氧化硅膜离心时,乙醇的加入会导致DNA结合。在洗涤去除污染物后,DNA用 200 μL 的 Tris-EDTA 溶液洗脱。
基因组 DNA 的预期产量将根据所用起始原料的数量和类型而有所不同(例如,不到一小时内,可从 2 x 106HeLa 细胞中分离出 15-25µg 经核糖核酸酶 A 处理的 DNA)。经试剂盒纯化的 DNA 的A260/A280比值在 1.6 和 1.9 之间,长度可达 50 kb。该 DNA 可用于限制性内切酶消化、PCR、Southern 印迹杂交和测序反应等下游应用。
需自备的试剂和设备
·55℃ 水浴或恒温水浴摇床
·70℃ 水浴或微量恒温仪
·带气溶胶屏障的移液器吸头(推荐)
·用于裂解的1.5 mL 微量离心管
·微量离心机(2 mL 管,装有转子)*
·乙醇 (95–100%),货号E7148或459836
·分子生物试剂水,货号W4502
*注:为了确保所有试剂管的正确放置,建议使用 24 位转子。如果您使用的是
36 位转子,我们建议您每隔一个位置妥善放置试剂管
储存方法和稳定性
室温储存试剂盒。如果任何试剂盒试剂形成沉淀物,请以 55–65°C 温热,直到沉淀物溶解,并在使用前冷却至室温。
制备说明
在开始该程序之前,执行以下操作:
1.将水浴或水浴摇床预热至 55°C
以使用组织、啮齿动物尾巴、新鲜全血和白细胞。
2.将水浴或微量恒温仪预热至 70°C
以使用培养细胞和组织。
3.彻底混匀试剂
检查试剂沉淀。如有任何试剂形成沉淀,以 55–65°C 温热直至沉淀溶解,并在使用前冷却至室温。
4.稀释洗液浓缩液
用 10 mL(10 份制备包)、80 mL(70 份制备包)或 360 mL(350 份制备包) 95-100% 乙醇稀释浓缩液。每次使用后,拧紧稀释洗涤液的瓶盖以防止乙醇挥发。
5.将蛋白酶 K 溶于水中
根据下表 1,将一瓶蛋白酶 K 粉末溶解于水中(货号W4502)以获得 20 mg/mL 的原液。此溶液可在 2–8°C 条件下保存数天。若要保存更长时间,溶液中未使用的部分使用前可在 –20°C 条件下分装保存。所提供的原样产品在室温下稳定。
每次必须将蛋白酶K溶液直接添加到每个样品中。切勿将蛋白酶 K 和裂解液合并存放。
操作流程
注意:如果在下游应用中需要最小剪切基因组 DNA,例如,如果使用最终产物进行长扩增 PCR,则通过温和移液或倒置混合至均匀,而不是在随后的程序中振荡。
A. 培养细胞制备
1a.收集细胞
• 贴壁细胞培养物:用胰蛋白酶释放细胞。以 300 ×g用5分钟离心沉淀5 × 106 个细胞;然后完全去除培养基并丢弃。继续步骤 2a。
• 悬浮细胞培养物:以 300 ×g用5分钟离心沉淀5 × 106 个细胞;然后完全去除培养基并丢弃。继续步骤 2a。
注意:细胞可收集并分装至 1.5 mL 微量离心管中,在液氮中快速冷冻,然后在
–70°C 条件下保存数月,至制备 DNA时取出。.
2a.重悬细胞
用 200 μL 重悬溶液彻底重悬细胞团块。如果事先冷冻,在重悬之前将细胞团块稍微解冻。如果不担心残留RNA,则直接前往执行步骤3a。
可选的RNase处理:如果需要无 RNA 的基因组 DNA,则可加入 20 μL 核糖核酸酶 A 溶液,室温水浴 2 分钟,然后继续步骤3a。
3a.裂解细胞
向样品中加入 20 μL 蛋白酶 K 溶液,接着加入 200 μL 裂解液 C (B8803)。彻底振荡(约 15 秒),然后70℃ 孵育 10分钟。均匀的混合物对于高效裂解是必不可少的。继续步骤 4。
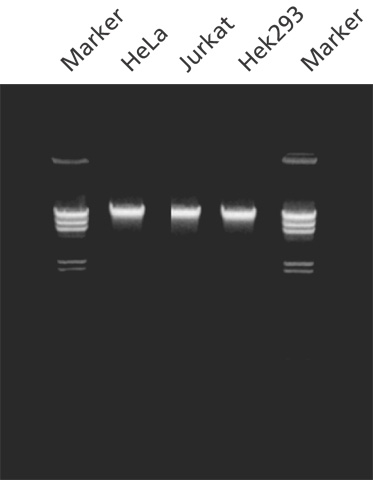
图 1.使用 GenElute™ 哺乳动物基因组 DNA 纯化试剂盒从细胞中纯化的基因组 DNA。用 GenElute™ 哺乳动物基因组 DNA 纯化试剂盒从指定来源的 2 x 106 个细胞中分离纯化的基因组 DNA。在 0.8% 琼脂糖凝胶上分析基因组DNA (200 ng/lane) 。标记物为用 Hind III 消化的 λDNA。
B. 哺乳动物组织制备
1b.制备组织
快速切片并称取一片新鲜或冷冻组织。切片前冰冻组织可稍稍解冻,但要放在冰上以防降解。将组织切成小块,以便更有效地裂解。每次制备可使用多达 25 mg组织(或 10 mg 脾脏,因为给定质量下的细胞量多)。转移至 1.5 mL 微量离心管中,继续步骤 2b。
注意:组织可收集并分装至 1.5 mL 微量离心管中,在液氮中快速冷冻,然后在
–70°C 条件下保存数月,至制备 DNA时取出。
2b.消化组织
向组织中加入 180μl 裂解液 T (B6678),然后加入 20μl 蛋白酶 K 溶液。涡旋混匀。55℃ 水浴样品,直至组织完全消化,无颗粒残留。偶尔振荡或使用水浴摇床。消化通常在 2~4 小时内完成。消化完成后,稍加振荡。
可选的RNase处理:如果不用担心残留RNA,可继续步骤 3b。如果需要无RNA的基因组DNA,则加入20 μL 核糖核酸酶 A溶液,在室温下孵育2分钟,然后前往执行步骤3b。
3b.裂解细胞
向样品中加入 200 μL 裂解液 C (B8803);彻底振荡(15 秒)。均匀的混合物对于高效裂解是必不可少的。在70 °C孵育10分钟。继续步骤 4。

图 2.使用 GenElute™ 哺乳动物基因组 DNA 纯化试剂盒从组织中纯化的基因组 DNA。用 GenElute™ 哺乳动物基因组 DNA 纯化试剂盒从指定小鼠组织中分离纯化的基因组 DNA。在 0.8% 琼脂糖凝胶上分析基因组 DNA (200 ng/lane) ,以说明产量和完整性。标记物为用 Hind III 消化的 λDNA。
C. 啮齿动物尾段制备
1c.制备啮齿动物尾段
快速测量并剪下一段新鲜或冷冻的啮齿动物尾巴。在切下前冷冻的啮齿动物尾巴可稍稍解冻,但要放在冰上以防降解。每次制备使用的尾段不要超过 0.6 cm(大鼠)或 1.2 cm(小鼠)。剪下两条(小鼠)或一条(大鼠)0.5-0.6 cm 长的尾巴,放入 1.5 mL 微量离心管中;继续步骤 2c。
注意:在制备 DNA 之前,啮齿动物尾巴可以在 –20°C 条件下贮存数月。
2c.消化组织
向鼠尾中加入 180μl 裂解液 T (B6678),然后加入 20μl 蛋白酶 K 溶液。振荡混匀;确保尾段完全浸没。55℃孵育样品,直至尾段组织完全消化。可能会残留一些颗粒(骨头和毛发)。培养过程中不时涡旋振荡或使用水浴摇床,以便更快速地消化。消化通常在 3~6 小时内完成。消化完成后,稍加振荡。如果不用担心残留RNA,可继续步骤 3b。
可选的RNase处理:如果需要无 RNA 的基因组 DNA,则可加入 20 μL 核糖核酸酶 A 溶液室温孵育 2 分钟,然后继续步骤3c。
3c.裂解细胞
向样品中加入 200 μL 裂解液 C (B8803);彻底振荡(15 秒)。均匀的混合物对于高效裂解是必不可少的。继续步骤 4。
D. 全血制备
1d.采集血液
采集全血置于抗凝管中(首选 EDTA 管)。全血在开始制备前应稳定至室温。
2d.血液制备
将 20 μL 蛋白酶 K 溶液加入 1.5 μL 微量离心管中。向试管中加入 200 μL 全血样品。如果样品小于 200 μL,加入重悬液使体积达到 200 μL。
注意:如果样品已经分装在试管中,可以将蛋白酶 K 溶液加入到样品中。涡旋振荡以确保酶充分混合。全血在制备 DNA 之前可在 4℃条件下保存数小时。如果不用担心残留RNA,可继续步骤 3d。
可选的RNase处理:如果需要无 RNA 的基因组 DNA,则可加入 20 μL 核糖核酸酶 A 溶液在室温下孵育2 分钟,然后继续步骤 3d。
3d.裂解细胞
向样品中加入 200 μL 裂解液 C (B8803);彻底振荡(15 秒)。均匀的混合物对于高效裂解是必不可少的。在55 °C孵育10分钟。继续步骤 4。
E. 白细胞 (WBC) 制备
1e.从全血中制备白细胞
每次从 500 μL 全血中制备 WBC;氯化铵裂解程序详见附录。
2e.重悬细胞
在 200 μL 重悬溶液中彻底重悬细胞团块;向样品中加入 20 μL 蛋白酶 K 溶液并稍加振荡,以确保酶充分混合。如果不用担心残留RNA,可继续步骤 3e。
可选的RNase处理:如果需要无 RNA 的基因组 DNA,则可加入 20 μL 核糖核酸酶 A 溶液,室温孵育 2 分钟,然后继续步骤 3e。
3e.裂解细胞
向样品中加入 200 μL 裂解液 C (B8803);彻底振荡。均匀的混合物对于高效裂解是必不可少的。在55 °C孵育10分钟。继续步骤 4。
从所有列出的样本类型中分离 DNA
这是以A、B、C、D 和 E 部分制备的样品继续进行的程序。
4.柱制备
向每个预装 GenElute™微量提取吸附柱(带红色 o 型环,不要与其他 GenElute™ 试剂盒混淆)中加入 500µL 柱制备液,并以12,000 × g 转速离心 1 分钟。弃去流出液。
注意:柱制备溶液可最大限度地提高DNA与膜的结合,从而获得更稳定的产量。
5.吸附准备
向裂解液中加入 200 μL 乙醇 (95–100%);涡旋振荡 5-10 秒以充分混合。溶液必须混合均匀。
6.加裂解液
从步骤 4 开始将管中的全部内容物转移到处理过的吸附柱中。在将内容物转移到吸附柱的过程中,可使用大口径移液器吸头以减少 DNA 剪切。以≥6500 × g 转速离心1分钟。丢弃含流出液的收集管,将吸附柱置于新的 2 mL 采集管中。
7.第一次洗涤
首次使用前,请按照配制说明书的说明用乙醇稀释洗涤浓缩液。向吸附柱中加入 500 μL 洗涤液,并以 ≥6,500 × g的转速离心 1 分钟。弃去含有流出液的收集管,将吸附柱置于新的 2 mL 收集管中。
8.第二次洗涤
向吸附柱中再加入 500 μl 洗涤液,并以最大速度(12,000-16,000 × g)离心3分钟干燥吸附柱。洗脱 DNA 前,吸附柱必须不含乙醇。如果看到残留乙醇,以最大速度将吸附柱再离心一分钟。如果您需要此额外的离心步骤,您可以清空并重新使用采集管。最后,丢弃含流出液的收集管,将吸附柱置于新的 2 mL 收集管中。
9.洗脱 DNA
将 200 μL 洗脱液直接移入吸附柱的中心;以 ≥6,500 × g 的转速离心 1 分钟以洗脱 DNA。为了提高洗脱效率,在加入洗脱液后在室温下孵育5分钟,然后离心。
可选: 重复步骤9进行第二次洗脱:用另外的 200 μL 洗脱液,洗脱到新的2 mL 收集管(已提供),或洗脱到第一洗脱所使用的 2 mL 收集管中。
洗脱液含有纯基因组DNA。对于 DNA 的短期贮存,建议使用 2-8°C。对于 DNA 的长期贮存,建议使用-20°C。避免冻融,以免引起DNA链断裂。洗脱液将有助于在这些温度下稳定 DNA。
DNA 沉淀(可选)
GenElute™ 哺乳动物基因组 DNA 试剂盒的设计使 DNA能够始终保持在溶液中,从而避免重悬问题。但是,如果您发现有必要浓缩 DNA,建议使用含乙酸钠的乙醇沉淀法。
结果
用 GenElute™ 试剂盒制备的基因组 DNA 的浓度和质量可以通过吸光度分析法和琼脂糖凝胶电泳法来确定。用 TE 缓冲液 (10 mM Tris-HCl, 1 mM EDTA, pH 8.0–8.5)稀释 DNA,用石英微量比色皿测量 260 nm 和 280 nm 处的吸光度。吸光度应介于 0.1 和 1.0 之间(或在分光光度计的线性范围内)。在 260 nm 处吸光度为 1.0 相当于约 50 μg/mL 的双链 DNA。260 nm 至 280 nm 处的吸光度比值 (A260/A280) 应为 1.6-1.9。DNA 的大小和质量可以通过琼脂糖凝胶电泳法来确定。1 在 0.5X TBE 缓冲液(货号T6400)中含有的 0.8% 琼脂糖的凝胶(货号A9539)可以很好地分离基因组 DNA。溴化乙锭(货号E1510)之类的嵌入染料可用来观察DNA,并对比已知的 DNA 标记物,例如 λ-DNAEcoRI Hind III 消化物(货号D9281),进行测量。基因组DNA应作为单一的高分子量条带迁移,几乎没有剪切迹象。脉冲场凝胶电泳法可以更精确地测定 DNA 大小。
故障排除指南
1. 吸附柱堵塞
原因— 样本太大。
解决方案— 之后使用更少的细胞、更小的组织块、更短的啮齿动物尾段或更少量的全血。若要挽救当前制剂,请提高转速和/或延长旋转时间,直到裂解液通过吸附柱。基因组 DNA 的产量可能降低。
原因—组织或啮齿动物尾段裂解效率低。
解决方案—延长蛋白酶 K 在 55°C 下的消化时间。为加速裂解,将组织或啮齿动物尾巴切成更小块,并在消化过程中频繁混合,以确保更高效的裂解。蛋白酶 K 消化后倒置样品管可确保混合更加均匀。切勿使用保存的合并溶液(蛋白酶 K+ 裂解液)。
2.基因组 DNA 产量低
原因—样本可能不新鲜或已降解,或组织裂解效率低。
解决方案—不同类型的细胞、组织和新鲜全血的产率不同。在培养物达到最大密度或完全融合之前使用它们。尽快收集组织或啮齿动物尾巴。在几小时内使用全血。如果样品要保存以供将来使用,请将细胞或组织在液氮中快速冷冻。全血在 4℃ 下保存时间不应超过 12 小时。
原因—裂解液/乙醇混合物不均匀。
解决方案—为确保溶液混匀,在应用到吸附柱前振荡 5–10 秒。如果在下游应用中需要最小剪切基因组 DNA,例如,如果使用最终产物进行长扩增 PCR,则通过温和移液或倒置混合至均匀,而不进行振荡。
原因—裂解液/乙醇混合物不均匀。
解决方案—为确保溶液混匀,在应用到吸附柱前振荡 5–10 秒。如果在下游应用中需要最小剪切基因组 DNA,例如,如果使用最终产物进行长扩增 PCR,则通过温和移液或倒置混合至均匀,而不进行振荡。
原因—DNA 洗脱不完全。
解决方案—确认 DNA 是用 200 μL 洗脱液洗脱的。将洗脱液加入吸附柱后在室温下孵育 5 分钟,大多数类型材料的 DNA 产量都会有所提高。可以使用 200 μL 洗脱溶液进行第二次和第三次洗脱。
原因—在吸附过程中漏用了乙醇。
解决方案—在步骤 6 中将样品用于吸附柱之前,检查步骤 5 中是否添加了乙醇。
原因—洗脱液中含有洗涤液中残留的乙醇。
解决方案—在洗脱 DNA 之前,必须去除最后洗涤液中残留的乙醇。离心更长时间或第二次干燥膜。如果含有乙醇流出液接触吸附柱,需在洗脱 DNA 之前重复离心步骤。
原因—洗脱液中含有洗涤液中残留的乙醇。
解决方案—在洗脱 DNA 之前,必须去除最后洗涤液中残留的乙醇。离心更长时间或第二次干燥膜。如果含有乙醇流出液接触吸附柱,需在洗脱 DNA 之前重复离心步骤。
原因—洗脱液中含有洗涤液中残留的乙醇。
解决方案—在洗脱 DNA 之前,必须去除最后洗涤液中残留的乙醇。离心更长时间或第二次干燥膜。如果含有乙醇流出液接触吸附柱,需在洗脱 DNA 之前重复离心步骤。
原因—洗涤浓缩液在使用前未经稀释。
解决方案—使用前检查洗涤浓缩液是否用乙醇适当稀释。
原因—洗涤浓缩液在使用前未经稀释。
解决方案—使用前检查洗涤浓缩液是否用乙醇适当稀释。
原因— DNA 用水而不是洗脱液洗脱。
解决方案—建议使用洗脱液洗脱,以实现纯化 DNA 的最佳产率和贮存。如果用水洗脱 DNA,请确认 pH 值至少为 7.0,以避免在酸性条件下储存较长时间后DNA会被酸水解。
3.DNA 的纯度低于预期;A260/A280比值太低
原因—样品被水稀释。
解决方案—使用洗脱液 (10 mM Tris-HCl,0.5 mM EDTA,pH 9.0) 或 10 mM Tris-HCl,pH 8.0–8.5 溶液作为洗脱剂。
原因—硅粉造成背景读数较高。
解决方案—将 DNA 样品以最大速度离心 1 分钟;使用上清液重复读取吸光度。
4.DNA 的纯度低于预期;A260/A280比值太高。
原因—基因组 DNA 被 RNA 污染。
解决方案—在之后的分离中加入核糖核酸酶 A 处理步骤,或用核糖核酸酶 A 溶液处理最终产物并进行再纯化。
5.DNA 被剪切
原因—基因组 DNA 处理不当。
解决方案—本试剂盒的设计可消除其他基因组 DNA 试剂盒中发现的可导致 DNA 剪切的 DNA 沉淀和常见重悬步骤。所有移液步骤应尽可能温和地执行。建议使用大口径移液器吸头,以帮助消除剪切。如果在下游应用中需要最小剪切基因组 DNA,例如,如果使用最终产物进行长扩增 PCR,则通过温和移液或倒置混合至均匀,而不进行振荡。
原因——样品陈旧、降解或已经历多次冻/融循环。
解决方案—旧的起始材料可能在洗脱液中产生降解DNA。应立即使用新鲜细胞和组织、鼠尾及全血制剂。或者,细胞和组织可以在液氮中冷冻并保存在-70°C 条件下,直至需要时取出。鼠尾可在-20℃条件下保存数周或-70℃ 条件下保存数月。全血可在 4℃ 条件下保存长达 12 小时。
6.下游应用被抑制
原因—乙醇被带入最终的基因组 DNA 制剂中。
解决方案—在最后一次洗涤吸附柱后(步骤 8),不要让洗涤液接触吸附柱。如有必要,在清空收集管后,再以最大速度(12,000–16,000 × g)离心柱子1分钟。
原因—盐被带入最终的基因组 DNA 制剂中。
解决方案—在步骤 8 中添加洗涤液之前,确保将吸附柱转移到新的 2 mL 收集管中。
附录: 从全血中分离白细胞的氯化铵裂解程序
1.在抗凝管中采集全血。
2.向 1 mL 氯化铵裂解液中加入 500 μL 全血*。在室温下轻轻混合(倒置或在摇床上)5 分钟,然后在微量离心机中以 700 × g 的转速离心 5 分钟。
3.弃上清液。将细胞团块轻柔重悬于另一毫升氯化铵裂解液中;按照步骤 2 温和混合并离心。弃去上清液,细胞团块置于冰上并重新开始白细胞程序 (1e)。
*氯化铵裂解液:
160 mM 氯化铵(货号A9434),20 mM 碳酸氢钠(货号S7277);用盐酸将 pH 调至 7.2。
材料
注意事项和免责声明
GenElute™ 哺乳动物基因组 DNA 试剂盒仅供实验室使用,不得用于药物、家庭或其他用途。裂解液 C 中含有离液盐,是一种刺激物。柱制备液具有刺激性。避免接触皮肤。处理这些溶液或随试剂盒提供的任何试剂时,请戴上手套、安全眼镜和合适的防护服。有关危险和安全处理方法的信息,请参阅安全数据说明书。
参考文献
如要继续阅读,请登录或创建帐户。
暂无帐户?