Peptide Library Design Tool Instructions
Welcome to the Peptide Library Design & Calculator tool presented
You may launch the Design Tool and view these instructions at the same time, or view only the reference charts.
Design overlapping peptide fragment libraries for your native protein of interest. Features of this peptide library design tool includes the ability to set custom sequence parameters (peptide length and amino acid offset), format the sequence output and export the custom peptide library sequences to other programs (Microsoft Word, Excel, etc).
- Enter the design tool.
- Select library type you wish to design.

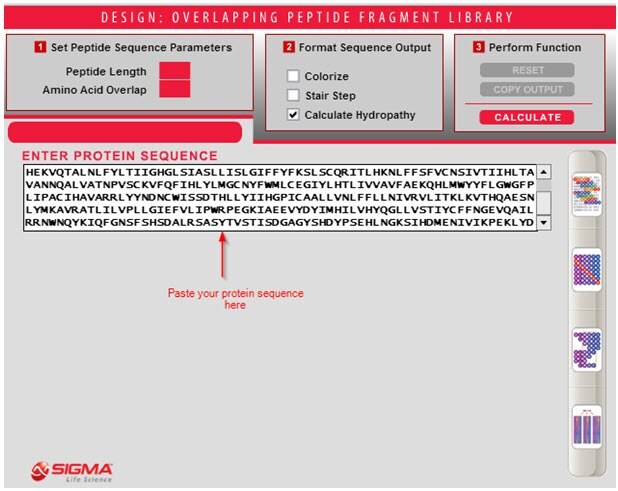
- Copy and paste the protein sequence of interest in the large text box labeled "ENTER PROTEIN SEQUENCE." This field accepts the 1-letter amino acid code, and accepts both upper and lower-case letters. Alternatively, you can type the protein sequence of interest in this field.

- Enter the desired peptide length. This peptide sequence length consideration will contribute to the success of the peptide library design. A value from 6 to 20 amino acids should be chosen for this field.
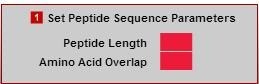
- Amino Acid Overlap: This value (from 1-20) determines the frame shift, or number of residues that the peptide sequence is shifted along the native protein sequence. A smaller overlap (i.e. 1 or 2) will ensure more chances for multiple epitope hits. A larger overlap (i.e. >6+) will result in a library with fewer peptides, however, presents the least chance for multiple epitope hits.
Peptide length and amino acid overlap are important when designing a peptide library, below is a table indicating the relationship between peptide length and amino acid offset number:
- Formatting peptide output. This section allows output customization of the peptide library sequences.
a) Colorize: This option will visually display, by color representation, the hydrophobic classification of each amino acid within the peptide sequence. This allows you to visually determine potential synthesis and/or solubilization difficulties of each peptide sequence, based upon the amino acid composition.

b) Stair Step: This option displays the peptide library sequences by the amino acid overlap as determined in the Sequence Parameters. Potentially missed epitopes can be quickly determined based on this format, as you are able to easily see the amino acid overlap relative to neighboring sequences. Example Protein Sequence: AVHNDTMTFRAPHSTWSEISILLQEVGSLGLSGDLRCGQRSQESP Calculated peptide sequences based on a peptide length of 15 and amino acid overlap of 5:

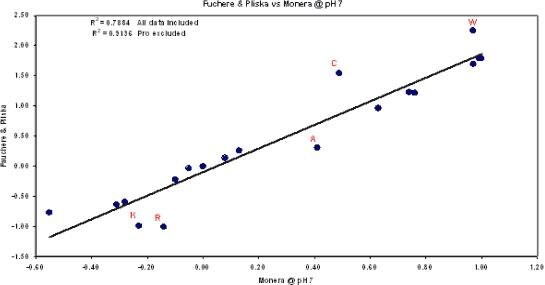
c) Calculate Hydropathy: This numerical value predicts the solubility characteristic of the peptide sequence in aqueous solutions. The value presented is based upon the weighted average known hydropathy values per amino acid divided by the length of the peptide sequence. The values are based on model systems of free energy transfer of acetyl-amino acyl amides (Ac-AA-am) by Fauchere & Pliska (1988) and based on the RP-HPLC retention times of peptide model systems at pH 7 by Monera.
Correlations between Fauchere & Pliska and Monera et al Hydrophobicity Scales are shown below:
- Click "CALCULATE" to generate the peptide library sequences based on the defined Sequence Parameters and Sequence Output. The total number of sequences generated will be shown at the bottom right of the screen.

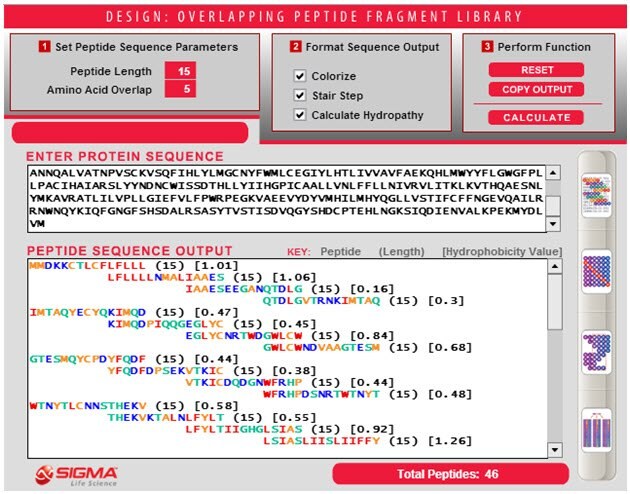
- Click "COPY OUTPUT" to export the sequences to another file (i.e. Microsoft Word, Excel, etc). The sequence format is automatically tab-delimited so all information will be placed as separate columns. After selecting "COPY OUTPUT," data is automatically copied and ready for pasting into your file type of choice. You will not be prompted that the sequence are ready for pasting.
To place an order using the output of the design tool, visit the PEPscreen Order Center.
Click "RESET" to clear all protein sequence and design results from the tool. This will reset (clear) all data so that you can begin a new design or modify the Peptide Sequence Parameters and/or the Format Sequence Output parameters.
If you have any questions or suggestions regarding this tool, please contact peptides@sial.com.
如要继续阅读,请登录或创建帐户。
暂无帐户?